Panning & Selection
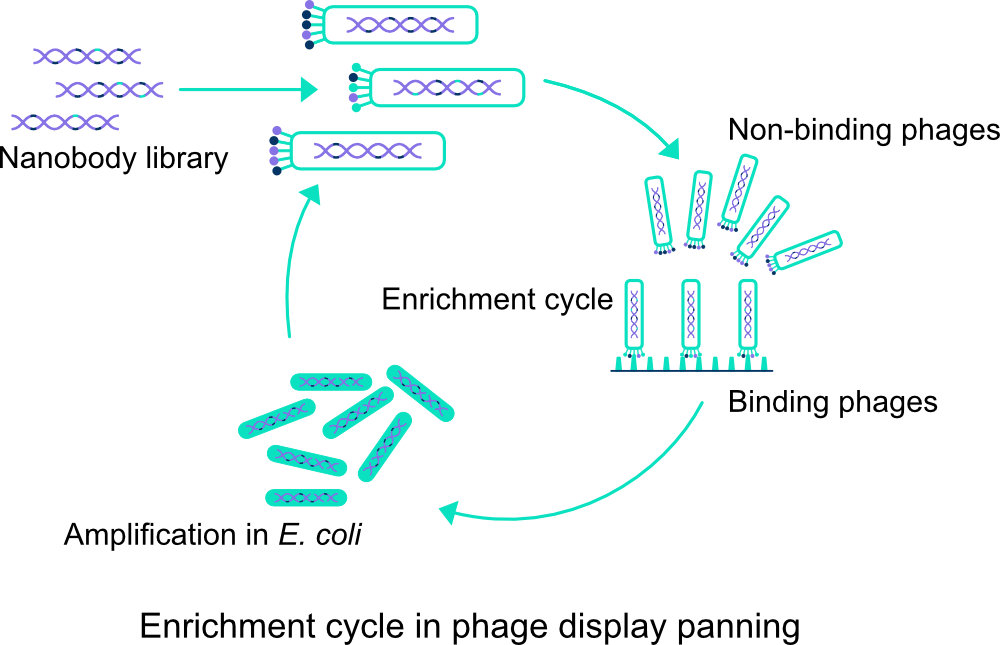
Our synthetic libraries have a great diversity and they have their own backbone structure in which only (parts of) the CDR regions are made variable. The fixed and variable DNA fragments of these nanobodies have been cloned into plasmids that have subsequently been incorporated into suitable E. coli strains. Using M13 phages, we can then express the nanobody in question and then screen and select it in various ways. The most ideal option is to ultimately screen for targets that are identical to the intended application of the nanobody.
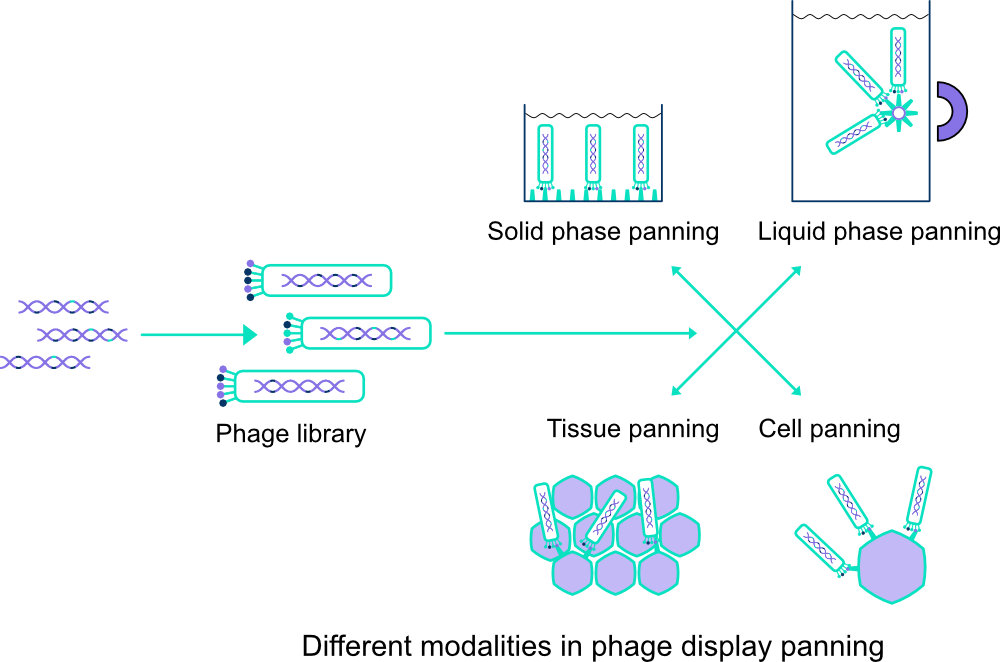
There are at least 4 ways in which we can select positive clones from the library using panning. The target of interest can be immobilized on, for example, a matrix such as wells in an ELISA plate (solid phase panning) or on metal particles in solution that can be kept out of solution with the aid of a magnet (liquid phase panning). We can also have clones bind to cells, attached or in solution, or to pieces of tissue with an even more natural structure. Finally, it may be useful to use combinations of techniques. Through a number of panning rounds, with washing steps in between, we obtain a selection of well-binding clones from which we will choose the best specimens. With DNA sequence analysis we gain more insight into the nanobodies and we can see how diverse the clones are.
Your content goes here. Edit or remove this text inline or in the module Content settings. You can also style every aspect of this content in the module Design settings and even apply custom CSS to this text in the module Advanced settings.
